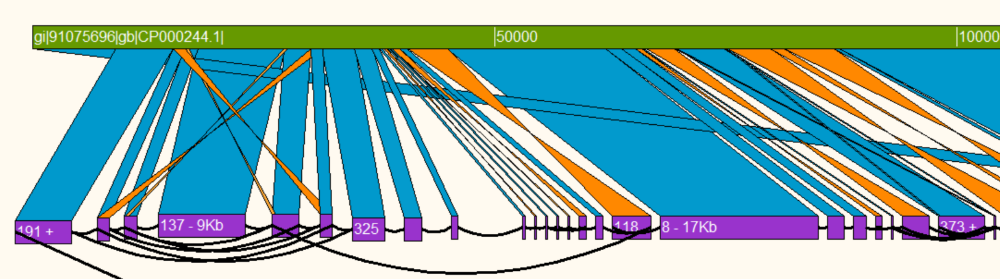
Contiguity
Contiguity is interactive software for the visualization and manipulation of de novo genome assemblies. Contiguity creates and displays information on contig adjacency which is contextualized by the simultaneous display of a comparison between assembled contigs and reference sequence. It is also useful in guiding manual closure of long read sequence assemblies. Contiguity is an open source application, implemented using Python and the Tkinter GUI package that can run on any Unix, OSX and Windows operating system. It has been designed and optimized for bacterial assemblies.
Contiguity was developed at the Beatson Microbial Genomics Lab.
For installation instructions, manual, example files and binaries go to Downloads.
For a quick start guide and more in depth tutorials go to Tutorials.
For a video overview of Contiguity and video tutorials visit Videos.
For information on how to cite Contiguity visit Citing.
If you have any problems with Contiguity or and feature suggestions post here.
I will endeavour to get back to you as soon as possible.
Email: mjsull@gmail.com
Follow @mjsull1
Updates
-
New release.
-
Updated website.